chapter 3 proteins protein structure (pages 125- 148; figures 3-1 to 3-28, 3-35) 1. the shape and structure of proteins - primary,
Chapter 3 Proteins
Protein structure (pages 125- 148; figures 3-1 to 3-28, 3-35)
1. The Shape and structure of proteins
- primary, secondary, tertiary, quaternary structure of proteins
- primary structure – sequence of amino acids; peptide bond
- secondary structures – -helix and -sheet; hydrogen bonds
- tertiary structure – noncovalent bonds; folding of proteins into a
conformation of lowest energy
- quaternary structure – noncovalent bonds
2. Protein domain arrangements (module)
- “In-line” – e.g. fibronectin type I, immunoglobulin
- “plug-in” – e.g. SH2 domain, kringle
3. Proteins can be classified into many families
4. Sequence searches can identify close relatives
- Generally a 30% identity suggests relatedness
5. Domain shuffling
6. Quaternary structure of proteins
- Weak bonds
- “head to head” arrangement – dimmers
- Single binding site
- “head to tail” arrangement – multimers
- 2 binding sites
- Ring – neuraminidase
- Filaments – actin
7. Proteins that have elongated, fibrous shapes
- Fibrous proteins
- e.g. alpha-keratin – intracellular
- collagen – extracellular
- Elastic fibers
- e.g. elastin – extracellular
8. Disulfide bonds stabilize extracellular proteins
- form in the ER
Protein function (pages 152-178; figures 3-36 to 3-67)
9. Selective binding of proteins to other molecules
- Binding may be weak or tight
- Specificity
- Weak bonds – ionic (electrostatic), hydrogen, van der Waals,
hydrophobic
- Binding site
10. Surface conformation of a protein determines its chemistry
- Interaction of neighboring parts of the polypeptide chain may
restrict the access of
water molecules to the protein’s binding site
*
Clustering of neighboring polar amino acid side chains can alter
their reactivity e.g. clustering of negatively charged side chains
increases affinity of a positively charged ion
11. The equilibrium constant measures binding strength
12. cAMP binding proteins – brings about conformational changes
- DNA binding proteins
- Enzymes (e.g. PKA)
- Ion channels
13. Serine proteases – “catalytic triad” – chemistry at an active site
14. SH2 domain – example of conserved binding sites
- Protein protein interactions – surface string, helix-helix,
surface-surface
15. Antibody molecules – specificity and affinity of binding sites
16. Equilibrium constant (K); Vmax; Km
- Turnover number = Vmax/enzyme concentration
17. Transition state; catalytic antibodies
- Stabilization of a transition state by an antibody creates an enzyme
18. Lysozyme – example of acid-base catalysis
- Distortion of bound substrate
- Negatively charged Asp attacks the C1 of the distorted sugar; Glu
donates a proton to the oxygen in the glycosidic bond; water molecule
displaces the Asp
19. General strategies for enzyme catalysis
*
Carbamoyl phosphate synthetase - molecular tunnels that connect
active sites
Glutamine NH3 carboxyphosphate carbamate carbamoyl phosphate
*
Pyruvate dehydrogenase complex – multienzyme complex
*
Aspartate transcarbamoylase – allosteric transition
*
Protein kinases/protein phosphatases – protein phosphorylation
*
Cdk – integrating protein
 TECNICA ANALITICA PARA LA DETERMINACION DE CLORUROS 1 PRINCIPIO
TECNICA ANALITICA PARA LA DETERMINACION DE CLORUROS 1 PRINCIPIO SALAT HABIENDO HECHO TUS ABLUCIONES Y CON LA FIRME
SALAT HABIENDO HECHO TUS ABLUCIONES Y CON LA FIRME FORMATO CONCEPTO DE LA SUBCOMISIÓN DE INVESTIGACIÓN SOBRE CALIDAD
FORMATO CONCEPTO DE LA SUBCOMISIÓN DE INVESTIGACIÓN SOBRE CALIDAD DZIAŁALNOŚĆ GOSPODARCZĄ NAJLEPIEJ ZACZĄĆ OD PODJĘCIA STRATEGICZNYCH DECYZJI DOTYCZĄCYCH
DZIAŁALNOŚĆ GOSPODARCZĄ NAJLEPIEJ ZACZĄĆ OD PODJĘCIA STRATEGICZNYCH DECYZJI DOTYCZĄCYCH 3084
3084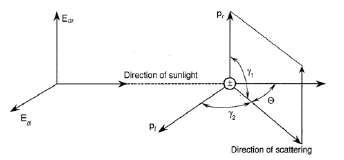 ATSC 5003 ATMOSPHERIC RADIATION LAB RAYLEIGH SCATTERING PHASE FUNCTION
ATSC 5003 ATMOSPHERIC RADIATION LAB RAYLEIGH SCATTERING PHASE FUNCTION PRESSEMELDING 090101 LIKESTILLINGSVEILEDER FOR KOMMUNENE LIKESTILLINGSSENTERET LEGGER TIRSDAG DEN
PRESSEMELDING 090101 LIKESTILLINGSVEILEDER FOR KOMMUNENE LIKESTILLINGSSENTERET LEGGER TIRSDAG DEN DIJITAL İNTERNET VE MOBIL BANKACILIK İSTATISTIKLERI12 EYLÜL 2018 BU
DIJITAL İNTERNET VE MOBIL BANKACILIK İSTATISTIKLERI12 EYLÜL 2018 BU TEUTH Y THAMUS VOZ Y ESCRITURA CERRAR “PUES BIEN
TEUTH Y THAMUS VOZ Y ESCRITURA CERRAR “PUES BIEN